 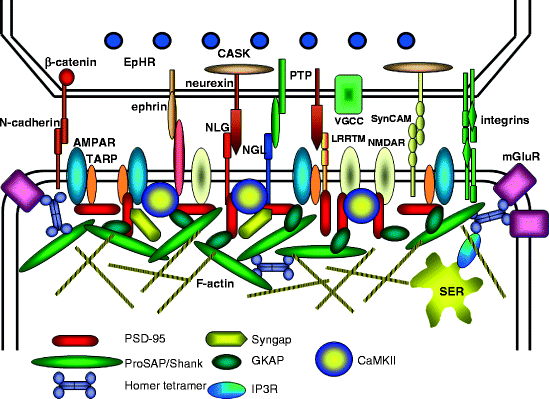
Motif RMSD AF versus native): dife_inp_1 (92 /0.65 Å), EFhand_inp1 (84,ĭesigns containing target-binding interfaces built around Stabilization of the protein upon binding calcium. Incubated with (4X protein concentration) and without CaCl 2 suggest (G) Tryptophan-enhanced terbium fluorescence spectra ofĪnd suggests the design can bind terbium. Purple, input non-motif residues in green, and overlaid with the native motifįrom 1PRW (orange). (F) AF2 prediction of inpainted designĮFhand_inp_1 scaffolding the double EF-hand motif with input motif residues in (protein concentration in CD experiments = 6.7 μM, Co 2+Ĭoncentration = 53.3 μM). Note that the coordination ofĬo 2+ in the protein core significantly stabilizes dife_inp_1 Both spectra areĬonsistent with the predicted helical structure. (D) CD spectra ofĭesign in the presence and absence of Co 2+. Using a predicted extinction coefficient of 155 for Co 2+ binding theīinding sites being recapitulated in the dife_inp_1 design. Quantification of the absorbance at 550 nm, (C) Titration analysis of Co 2+ against the design (proteinĬoncentration = 200 μM). Sequence but with the 6 coordinating residues (sidechains shown in (A)) mutated Note the peaks at 520 nm, 555 nm andĦ00 nm, consistent with Co 2+ binding to the desired scaffolded motif (B)Ībsorbance spectra showing of dife_inp_1 (or mutant) in the presence (or not) ofĪn 8-fold molar excess of Co 2+. Hallucinated scaffold (gray) hallucinated motif (purple) bound metal (blue).Īctive site residues shown in boxes for di-iron and EF-hand respectively. Native protein scaffold (light yellow) native functional motif (orange) Wavelength, from CD spectroscopy, for the 3 RSV-F site V hallucinations with (D) Mean residue ellipticity (MRE) versus K D values forĮach design are shown in legend. RSV-F refers to purified trimeric native F protein. Point mutants at various concentrations of hRSV90 antibody, with sigmoid fits. Maximum SPR signal (response units) of purified RSV-F epitope scaffolds and See Table S2 for additional metrics on each design. Motif (orange) hallucinated scaffold (gray) hallucinated motif (purple) īinding partner (blue). Colors: native protein scaffold (light yellow) native functional Interface helix (6VW1 chain A residues 24–42). (B) Design of COVID-19 receptor trap based on ACE2 For RSV-F designs, these metrics are rsvf_ii_141 (85.0, 0.53 The accuracy of recapitulation of the original scaffolded motif (motif RMSD AF Metrics computed on the AF2 predictions: prediction confidence (AF pLDDT), and Sequences which fold to structures harboring the desired motif through two Here and in the following figures, we assess the extent of success in designing S9 here because of spaceĬonstraints we show only the AF2 model the two are very close in all cases. Structure predictions from the design sequence are in Fig. Comparisons of the RF hallucinated models to AF2 (site II: PDB 3IXT chain P residues 254–277 site V: 5TPN chain A (A) Design of proteins scaffolding immunogenic epitopes on RSV protein F 
Masked region (light gray, dotted lines). Native functional motif (orange) hallucinated/inpainted scaffold (gray) Ĭonstrained motif (purple) binding partner (blue) non-masked region (green) Circles: Average of 20 outputsįor each of the benchmarking proteins. “motif”) to the original structure (F), and in AlphaFoldĬonfidence (pLDDT in the replaced region) (G). Hallucination, both in terms of the RMSD of the unmasked protein (the Length, RF joint typically modestly outperforms Were tasked with filling in the missing sequence and structure to For each protein, 20 random masks of lengthģ0 were generated, and RF joint and hallucination Set of 28 de novo designed proteins, published since (F-G) Motif scaffolding benchmarking data comparing Resemble the original protein (2KL8, left) and are confidently predicted byĪlphaFold (pLDDT/Motif RMSD of models shown: 91.6/0.91, 92.0/0.69, 90.4/0.82 Outputs (inpainted region in gray) closely Of sequence and structure masked out, with the network tasked with predicting RF joint with a continuous (length 30) window Structure and sequence of a masked region of protein. (D) Protein design challenges formulated as missing Information is input into a modified RosettaFold network (termed Recapitulation and other task-specific functions. S2) which are scored by a lossįunction that rewards certainty of the predicted structure along with motif Residue-residue distances and orientations (Fig. TrRosetta or RosettaFold neural network, which predicts 3D coordinates and At each iteration, a sequence is passed to the (A) Applications of functional-site scaffolding. 
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
